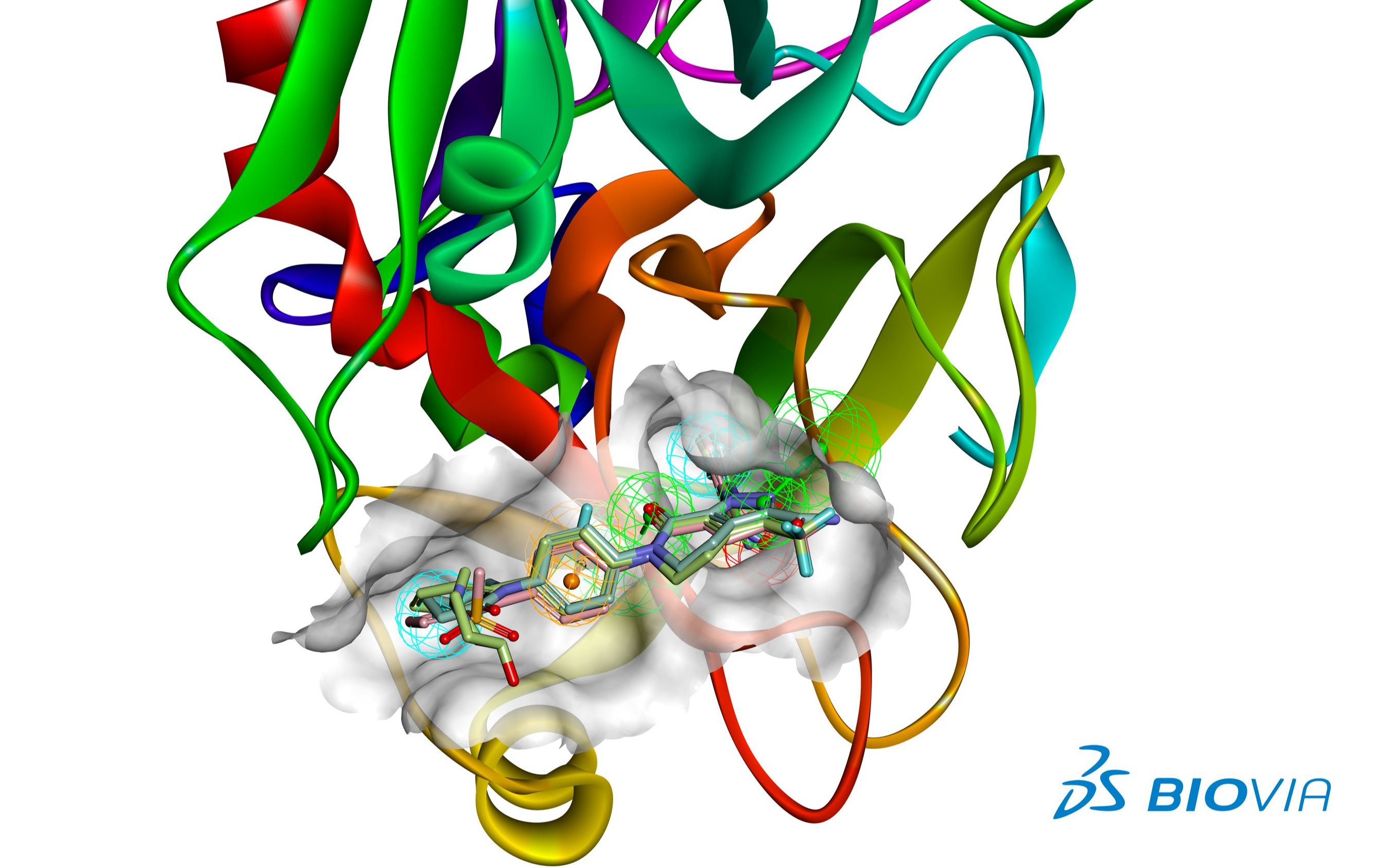
Structure alignment is a valuable tool for the comparison of proteins with low sequence similarity, where evolutionary relationships between proteins cannot be easily detected by standard sequence alignment techniques. One such example is circular permutation, where the relative locations of structural elements (and the N- and C-termini) within two proteins are different, but their overall shape and structure (e.g., secondary structural elements and their relative orientations) are conserved. There are many examples of natural and designed proteins where the spatial arrangement of secondary structural elements or protein domains is maintained but the protein backbone connections between these structural elements are different - i.e., the proteins have different topologies. However, this assumption may not always be true. Most structure alignment algorithms assume that the structural units of two similar proteins appear in the same order (in the N-terminal to C-terminal direction) within their sequences. For example one of these proteins may contain extra loops or truncations that alter relative orientation of different domains in the structures.
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
